Métabolismes énergétiques microbiens et enzymes redox
Microbial energy metabolisms and redox enzymesResearch (En)
Our group studies various bacterial redox enzymes, from their molecular mechanisms to their physiological roles. The aim is to gain a better understanding of how these enzymes work and of the energy metabolism of microorganisms.
Most of these enzymes are involved in the energy metabolism of hydrogen, sulfur, and oxygen. More recently, the team has focused on enzymes and metabolic pathways that use or release CO2. In addition to their molecular characterization (using various biochemical or biophysical approaches: purification of soluble or membrane proteins or complexes, enzymatic activities, native blue gels, spectroscopy, electrochemistry, etc.), the aim is to determine the function of some of these enzymes in cellular energy metabolism. The role of these proteins in the cell is addressed by combining molecular biology, microbiology, genetics, and biochemistry, such as bacterial growth under different conditions, gene deletions, and the study of changes in the metabolome, transcriptome, and proteome. These approaches allow us to reconstruct the energy metabolic pathways we are studying.
Various bacteria are cultivated from a physiological and phylogenetic point of view: Solidesulfovibrio fructosivorans (sulfate-reducing anaerobe) and Aquifex aeolicus (hyperthermophile, microaerophile, hydrogen and sulfur-oxidizing). Enzymes from environmental or pathogenic bacteria are also studied: Escherichia coli, Helicobacter pylori, and Dehalococcoides mccartyi.
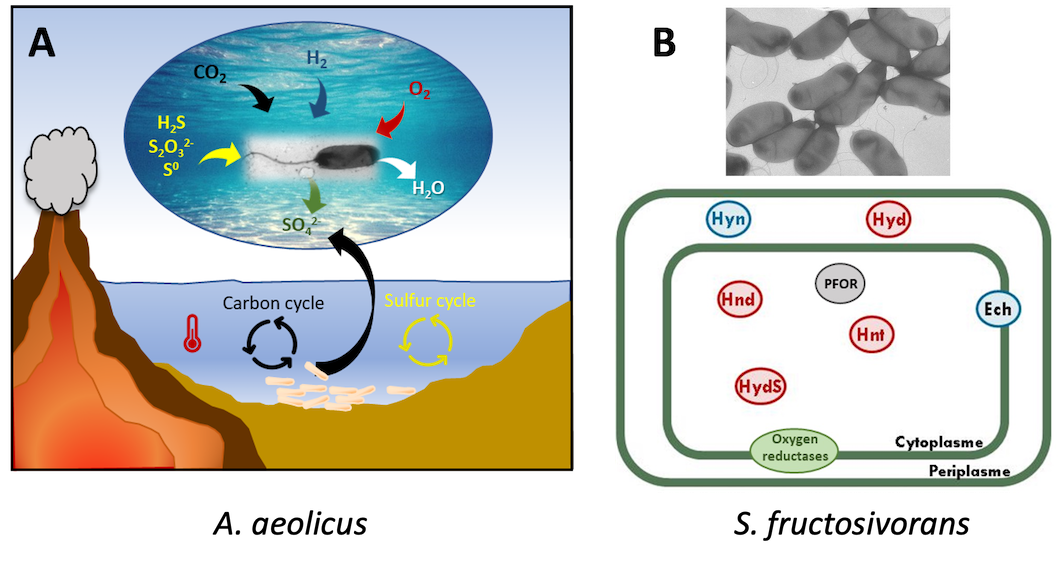
A: A. aeolicus is a marine bacterium isolated near shallow underwater volcanic vents on the island of Vulcano (Italy). It feeds on molecular hydrogen, carbon dioxide, molecular oxygen, and minerals, releasing only water. In addition to H2, inorganic sulfur compounds can also be oxidized to sulfate. (Guiral & Giudici-Orticoni, 2021). B: S. fructosivorans is a mesophilic anaerobic bacterium isolated from sediments. It has a complex system of hydrogenases.
Our research focuses primarily on hydrogenases, which are enzymes that reversibly catalyze the oxidation of molecular hydrogen into protons. Our work aims to study the role of hydrogenases in cellular energy metabolism and the reaction mechanisms of these enzymes. We are developing an integrated multidisciplinary approach to hydrogen metabolism, working closely with the unit’s teams of chemists and physicists, from the enzyme and the catalytic act to its cellular functions and its role in energy-generating mechanisms. Our work therefore falls within the laboratory’s “hydrogen and associated metabolisms” research area. We are studying several biological models, including the sulfate-reducing bacterium Solidesulfovibrio fructosivorans, which is particularly interesting because it has a complex system of six different hydrogenases: four [FeFe] hydrogenases and two [NiFe] hydrogenases. Why is there such diversity of hydrogenases in a single organism? What is the role of these enzymes?
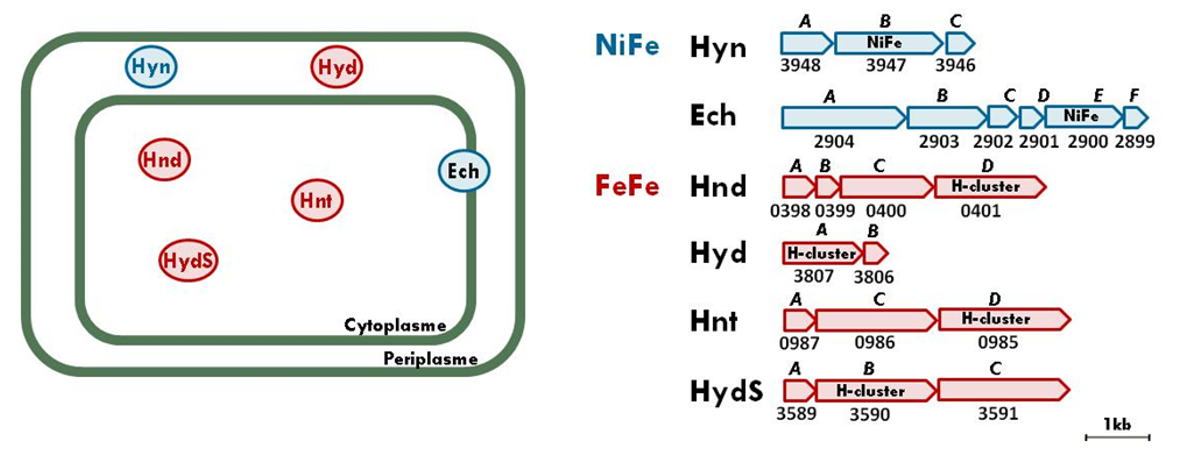
Solidesulfovibrio fructosivorans and its hydrogenase system: on the left, their cellular localization; on the right, their operons (Baffert et al., 2019)
In recent years, our research has focused on studying an [FeFe] hydrogenase from this organism, Hnd, which performs electron bifurcation. Enzymes that can bifurcate electrons perform the energy coupling of an exergonic redox reaction with an endergonic redox reaction.
It has been shown that these hydrogenases couple proton reduction with the endergonic oxidation of NADH and the concomitant exergonic oxidation of reduced ferredoxin (and/or the reverse reaction). We are studying this enzyme from its molecular functioning to its physiological role in bacteria. With regard to its involvement in bacterial physiology, we have shown that Hnd is involved in ethanol metabolism. It is a reversible hydrogenase that functions by consuming H2 through an electron bifurcation mechanism during pyruvate fermentation and by producing H2 through an electron confurcation mechanism when the bacterium uses ethanol as an electron donor, a condition in which Hnd is essential for bacterial growth.
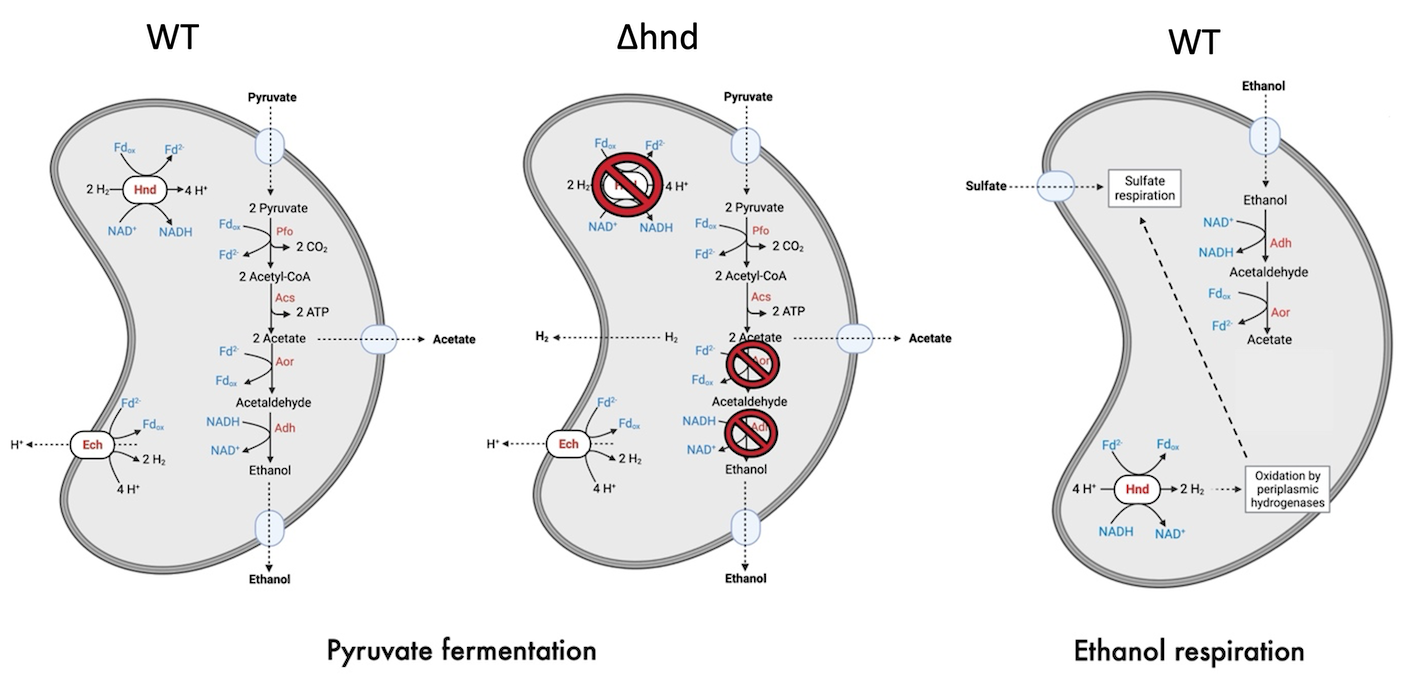
Pyruvate fermentation and ethanol oxidation pathways in the wild-type (WT) and mutant (∆hnd) strains of S. fructosivorans (Payne et al. 2022; Kpebe et al. 2023)
Our research also aims to remove the scientific barriers to the use of hydrogenases in biotechnological processes, and in particular to understand the molecular basis of hydrogenase inactivation by O2, which is one of the main limitations at present. This research is being conducted in collaboration with the BIP6 team. Dihydrogen is considered a “clean” fuel of the future, an alternative to fossil fuels, provided that it is no longer produced using fossil energy. In this context, the conversion of solar energy into H2 is an attractive strategy for the bioproduction of this fuel.
The ANR supports our research on hydrogenases: Engineering-H2 Cyano, 2009-2013; Algo-H2, 2011-2015; Cyanhy, 2013-2017; Heros, 2014-2019; Otolhyd, 2019-2023; Hydre, 2024-2028.
Within the team, we also conduct research on terminal oxidases and bacterial aerobic respiration. Currently, we are particularly interested in bd-type oxidase (from Solidesulfovibrio fructosivorans and Escherichia coli). Using biochemical, biophysical, and physicochemical approaches, in collaboration with the laboratory’s “Biophysics of Metalloproteins and Dynamic Systems” team, we are conducting a molecular and functional study of this enzyme.
The team is also seeking to understand the role of different terminal oxidases (bd-type or heme-copper oxidases) in the hyperthermophilic bacterium Aquifex aeolicus, which grows optimally in the presence of low concentrations of O2.
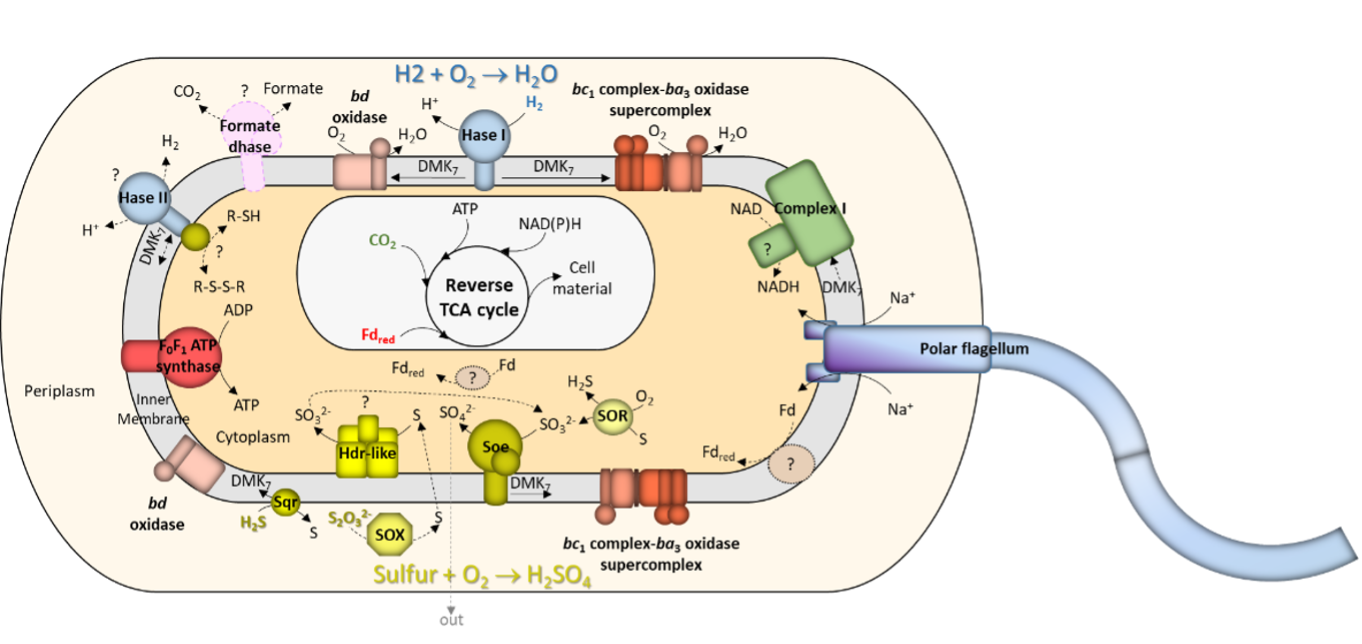
Energy conservation and carbon assimilation in Aquifex aeolicus (Guiral & Giudici-Orticoni, 2021)
The prokaryotic oxidative metabolism of sulfur is complex and not fully understood. We are attempting to identify and characterize enzymes involved in these oxidation pathways (e.g., sulfite dehydrogenase SoeABC, heterodisulfide reductase complex Hdr, and sulfur transferases) in the hyperthermophilic bacterium Aquifex aeolicus, using mainly biochemical and proteomic approaches.
This characterization of sulfur enzymes has recently led us to take an interest in the conversion of arsenic (reduced and oxidized forms) and thioarsenic (in collaboration with the BIP “Evolution of Bioenergetics” team), which could occur in certain bacteria through enzymes known to use sulfur compounds (such as the Sox system for oxidizing thiosulfate S2O32-). This aspect of the work involves molecular biology, microbiology, and biochemistry.
- Diversity of pyruvate ferredoxin oxidoreductases (PFOR)
Pyruvate ferredoxin oxidoreductase (PFOR) is a metalloenzyme that reversibly catalyzes the following reaction:
CoA + pyruvate + Fdox ⇄ Acetyl−coA + CO2 + Fdred (Fd = ferredoxin)
It is an essential enzyme in the central metabolism of various anaerobic pathogenic bacteria and parasites and is not present in mammals. It is therefore very interesting to find specific inhibitors of the active site of PFORs. These enzymes have been relatively little studied, so it is also interesting to better understand their reactivity, diversity, and structure-function relationships.
We produce and characterize PFOR from different organisms at the molecular level in order to determine their catalytic properties and sensitivity to inhibitors, including oxygen. The synthesis of new potential inhibitors is carried out in collaboration with Michel Maffei (ISm2, Marseille).
- CO2 fixation in A. aeolicus
We have begun studying certain aspects of CO2 fixation in A. aeolicus, which is obligate autotrophic. This bacterium assimilates CO2 via the reverse tricarboxylic acid cycle (rTCA), which involves PFOR and OGOR, two pentameric enzymes that use reduced ferredoxin to function, which we have characterized. The aim of this work is to explore the ability of this microorganism to couple CO2 fixation via the rTCA to the oxidation of hydrogen or sulfur compounds and to understand how the low redox potential ferredoxins involved in carbon fixation are reduced in A. aeolicus.
